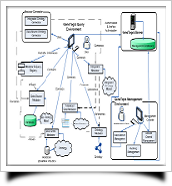
GeneTegra leverages Semantic Web technologies to facilitate the exploration, aggregation, and querying of the vast wealth of biomedical data that is constantly being generated and updated by investigators and data managers. Using current and emerging technology standards, GeneTegra delegates queries to the live data sources in a federated environment, ensuring data freshness and avoiding the need for costly storage of duplicate data.
GeneTegra makes use of the following technology standards:
- OWL (Web Ontology Language), a Semantic Web standard, is used for modeling of data sources and schemas.
- RDF (Resource Description Framework), a Semantic Web standard, is used for modeling of data instances.
- SPARQL, a Semantic Web standard, is used as the native query language of the system.
- WSDL (Web Services Description Language) is used for describing remote access mechanisms for data sources.
- JDBC (Java Data Base Connectivity) is used to connect the GeneTegra server to underlying SQL-based databases.
- SQL (Structured Query Language) is used to query relational databases.
- XML (eXtensible Markup Language) is used to store and depict configuration files, and is also a format for potential sources of data.
- HTTP (Hypertext Transfer Protocol) is used for remote access to data sources.
- Java is used as the software programming language and runs over the standard Java Virtual Machine, version 6 or higher.
